ELECTRON TRANSPORT CHAIN (ETC)
|
The
respiratory chain
is embedded in cristae of the inner membrane.
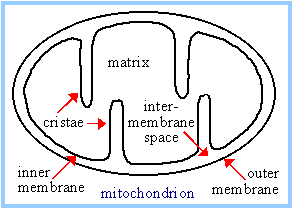
Spontaneous electron
transfer through respiratory chain complexes I, III, & IV is coupled to H+
ejection from the matrix to the intermembrane space. Because the outer
membrane contains large channels, protons in the intermembrane space
equilibrate with the cytosol. Respiration-linked pumping of protons out of
the mitochondrial matrix conserves some of the free energy of spontaneous
electron transfers as potential energy of an electrochemical H+
gradient.
Electron transfer from
NADH to
O2 involves
multi-subunit inner membrane
complexes I, II, III, & IV, plus
coenzyme Q and
cytochrome c. Within each
complex, electrons pass sequentially through a series of electron carriers.
The composition of each
of the respiratory chain complexes
is shown below:
|
Complex |
Name |
No. of Proteins |
Prosthetic Groups |
|
Complex I |
NADH Dehydrogenase |
46 |
FMN, 9 Fe-S centers |
|
Complex II |
Succinate-CoQ Reductase |
5 |
FAD, cyt b560,
3 Fe-S centers |
|
Complex III |
CoQ-cyt c Reductase |
11 |
cyt bH, cyt
bL, cyt c1, Fe-SRieske |
|
Complex IV |
Cytochrome Oxidase |
13 |
cyt a, cyt a3,
CuA, CuB |
|
|
|
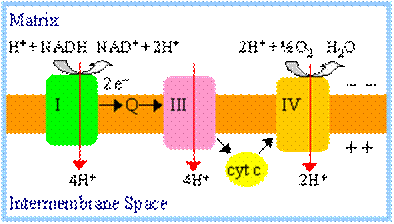
A total of 10 protons
are ejected from the mitochondrial matrix per 2 electrons transferred from
NADH to oxygen via the respiratory chain. The H+/e-
ratio for each respiratory chain complex will be discussed separately.
|
|
|
Complex I (NADH
Dehydrogenase) transports 4H+ out of the mitochondrial matrix per
2e- transferred from
NADH to coenzyme Q. Complex I catalyzes oxidation of NADH, with reduction
of coenzyme Q
NADH + H+
+ Q ® NAD+ + QH2
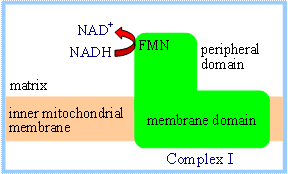
Lack of high-resolution structural
information for the membrane domain of complex I has hindered elucidation of
the mechanism of H+ transport through this complex. Direct
coupling of transmembrane proton flux and electron transfer is unlikely,
because the electron-transferring prosthetic groups, FMN and iron-sulfur
centers, are all located in the peripheral domain of complex I. Thus it is
assumed that protein conformational changes are involved in H+
transport, as with an
ion pump.
|
Succinate
Dehydrogenase of the
Krebs Cycle
is also called complex II
or Succinate-CoQ Reductase.
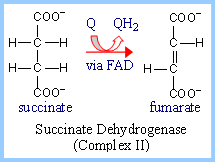
FAD is the initial electron receptor.
FAD is reduced to FADH2 during oxidation of succinate to
fumarate. FADH2 is then reoxidized by transfer of electrons
through a series of three iron-sulfur centers to Coenzyme Q, yielding QH2.
|
|
|
X-ray crystallographic
analysis of E. coli complex II indicates a linear arrangement of
electron carriers within complex II, consistent with the predicted
sequence of electron transfers:
FAD ® FeScenter 1
® FeScenter 2
® FeScenter 3
® CoQ
|
|
|
Complex III (bc1
complex): H+ transport in complex III involves coenzyme Q (CoQ).
The "Q cycle" depends on the mobility of CoQ within the lipid bilayer.
There is evidence for one-electron transfers, with an intermediate
semiquinone radical state.
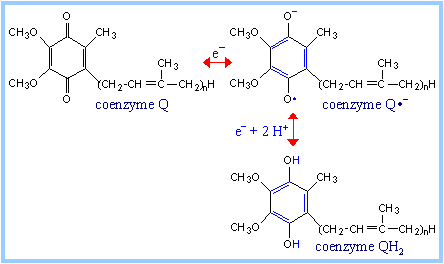
Q Cycle:
|
|
|
|
- Electrons enter complex
III via coenzyme QH2, which binds at a site on the positive
side of the inner mitochondrial membrane, adjacent to the intermembrane
space.
- QH2 gives up
one electron to the Rieske iron-sulfur center (Fe-S).
Fe-S is reoxidized by transfer of the electron to cytochrome c1, which
passes it out of the complex to cytochrome c.
The loss of one electron from QH2 would generate a semiquinone
radical, shown here as Q·-,
although the semiquinone might initially retain a proton as QH·.
- A second electron is
transferred from the semiquinone to cytochrome bL (heme bL),
which passes it across the membrane via cytochrome bH
(heme bH) to another CoQ bound at a site on the matrix
side of the membrane.
The fully oxidized CoQ, generated as the second electron is passed to the
b cytochromes, may then dissociate from its binding site adjacent to the
intermembrane space.
- Accompanying the
two-electron oxidation of bound QH2, 2H+ are
released to the intermembrane space.
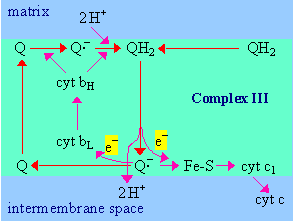
|
|
In an alternative mechanism that has been
proposed, the two electron transfers from QH2 to Fe-S & cyt bL
may be essentially simultaneous, eliminating the semiquinone intermediate.
It takes 2 cycles for CoQ, bound at the site near
the matrix side of the membrane, to be reduced to QH2, as it accepts
2 electrons from the b hemes and 2 H+ are extracted from the matrix
compartment. In 2 cycles, 2 QH2 enter the pathway, and one is
regenerated.
Overall reaction catalyzed by complex III, including net inputs and outputs
of the Q cycle:
QH2 + 2H+(matrix side) + 2 cyt c
(Fe3+) à Q + 4H+(outside)
+ 2 cyt c (Fe2+)
Per
2e-
transferred through the complex to cytochrome c,
4H+ are released to
the intermembrane space. While 4H+ appear outside per net 2e-
transferred in 2 cycles, only 2H+ are taken up on the matrix side. In
respiratory chain
complex IV
(see below), there is a similarly uncompensated uptake of protons from the
matrix side (4H+ per O2 or 2 per 2e-).
Thus there are 2H+ per 2e-
that are effectively transported by a combination of complexes III & IV. They
are listed with complex III in diagrams depicting H+/e-
stoichiometry.
|
The b hemes are positioned to provide a
pathway for electron transfer across the membrane.
The protein domain with attached Rieske
iron-sulfur center (labeled Fe-S) has a flexible link to the rest of the
complex. At right, the iron-sulfur center protein is colored green. The
iron-sulfur center changes position during electron transfer. After Fe-S
extracts an e- from QH2,
it moves closer to heme c1
(cytochrome c1) to which
it transfers the e-.
After the first electron transfer from QH2 to Fe-S, the CoQ
semiquinone is postulated to shift position within the Q-binding site,
moving closer to its electron acceptor, heme bL. This would help
to prevent transfer of the second electron from the semiquinone to Fe-S.
|
|
Complex
IV (Cytochrome Oxidase): Electrons are donated to complex IV,
one at a time, by cytochrome c, which binds from the intermembrane space. Each
electron passes via CuA and heme a to the
binuclear center,
buried within the complex, that catalyzes oxygen reduction:
4e-
+ 4H+ + O2 → 2H2O
Protons utilized in this reaction are taken up
from the matrix compartment.
In addition to the protons utilized in the
reduction of O2, there is electron transfer-linked transport of 2H+
per 2e- (4H+ per
4e-) from the matrix to the
intermembrane space.
Structural and mutational studies indicate that
protons pass through complex IV via chains of groups subject to protonation/deprotonation,
called "proton wires." These consist mainly of chains of buried water molecules,
along with amino acid side-chains, and propionate side-chains of the hemes.
Separate H+-conducting pathways link
each side of the membrane to the buried
binuclear center
where O2 reduction takes place. These include two proton pathways,
designated "D" and "K" (named after constituent Asp and Lys residues) extending
from the mitochondrial matrix to near the binuclear center deep within complex
IV.
A switch mechanism controlled by the reaction cycle is proposed to effect
transfer of a proton from one half-wire (half-channel) to the other. There
cannot be an open pathway for H+ completely through the membrane, or
oxidative phosphorylation would be
uncoupled.
(Pumped protons would leak back;). The process of switching may involve
conformational changes, and oxidation/reduction-linked changes in pKa
of groups associated with the catalytic metal centers.
|
The ATP synthase,
which is embedded in cristae of the inner mitochondrial membrane, includes
the following major subunits:
·
F1
- the catalytic subunit, made of 5 polypeptides with stoichiometry
a3b3gde.
·
Fo
- a complex of integral membrane proteins that mediates proton transport.
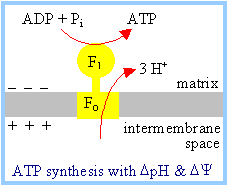
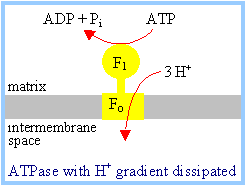
The F1Fo
complex couples ATP synthesis to H+ transport into the
mitochondrial matrix. Transport of at least 3H+ per ATP
synthesized is required, as estimated from a comparison of the following:
|
|
- the free energy change (DG)
associated with synthesis of ATP
under cellular conditions (the free energy required)
- the free energy change (DG)
associated with transport of each H+
into the mitochondrial matrix, based on the electrochemical H+
gradient (the free energy available per H+).
|
The
Chemiosmotic Theory of oxidative phosphorylation, for which Peter Mitchell
received the Nobel prize is summarized in the diagram.
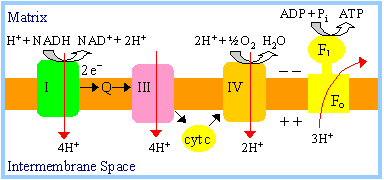
The Chemiosmotic
Theory states that coupling of electron transfer to ATP synthesis is
indirect, via a H+ electrochemical gradient: |
|
1.
Respiration:
Spontaneous electron transfer through complexes I, III, and IV is coupled to
non-spontaneous H+ ejection from the mitochondrial matrix. H+
ejection creates a membrane potential
(DY, negative in the matrix) and a
pH gradient (DpH,
alkaline in the matrix).
2.
F1Fo ATP
Synthase: Non-spontaneous ATP synthesis is coupled to
spontaneous H+ transport into the matrix compartment. The pH and
electrical gradients created by respiration are together the driving force for H+
uptake.
Return of protons to the matrix via Fo "uses up" the pH and
electrical gradients.
|
ATP produced in the
mitochondria must exit to the cytosol to be used by transport pumps,
kinases, etc. ADP
and Pi,
arising from ATP hydrolysis in the cytosol, must re-enter the mitochondria
to be converted again to ATP.
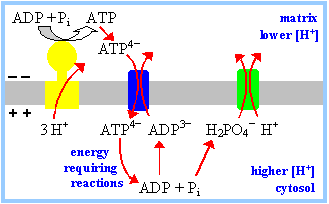
Two
carrier proteins in the inner mitochondrial membrane are required for
this metabolic cycle. The outer membrane is considered to be not a
permeability barrier. The large VDAC channels in the outer membrane are
assumed to allow passage of adenine nucleotides and Pi. |
|
1.
The Adenine Nucleotide Translocase (ADP/ATP carrier) is an antiporter
that catalyzes exchange of ADP for ATP across the inner mitochondrial membrane
(p. 496). At cellular pH, ATP has four negative charges, while ADP has 3
negative charges. ADP3-/ATP4-
exchange is driven by, and uses up, the membrane potential generated by
respiration (one charge per ATP).
2.
Phosphate reenters the mitochondrial matrix with H+, by an
electroneutral symport mechanism. Pi entry is driven by and uses up
the pH gradient (equivalent to one mole of H+ per mole of ATP).
|
Thus the equivalent of one mol of H+
enters the matrix with ADP/ATP exchange and Pi uptake. Assuming
transport of 3 mol H+ by F1Fo, a
total of 4H+
would enter the mitochondrial matrix
per ATP synthesized. |
The phenomenon of
respiratory control is the subject of important study. An oxygen electrode may
be used to record [O2] in a closed vessel. Electron transfer, e.g.,
from NADH to O2, is monitored by recording the rate of disappearance
of O2.
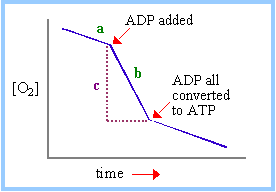
|
At right is an
idealized representation of an oxygen electrode recording while
mitochondria respire in the presence of Pi, along with an
electron donor (e.g., succinate, or a substrate of a reaction that will
generate NADH).
The dependence of
respiration rate on availability of ADP, the substrate for the ATP
Synthase, is called respiratory control. The respiratory control ratio is
the ratio of slopes after and before ADP addition (b/a). The P/O ratio is
the moles of ADP added, divided by the moles of O consumed (based on c)
while phosphorylating the added ADP. |
|
Chemiosmotic explanation of respiratory
control:
Electron transfer is obligatorily coupled to H+
ejection from the matrix. Whether this coupled reaction is spontaneous depends
on the pH and electrical gradients.
|
Reaction |
Free energy change |
|
e-
transfer (e.g., NADH to O2) |
a negative value* |
|
H+ ejection
from the matrix |
a positive value that
varies with the H+ gradient** |
|
e-
transfer coupled to H+ ejection |
algebraic sum of the
above |
*DGo'
= - nFDEo'
= -218 kJ/mol, for transfer of 2 e-
from NADH to O2.
** For ejection of one H+ from the
matrix:
DG = RT ln ([H+]cytosol/[H+]matrix)
+ F DY = 2.3 RT (pHmatrix
- pHcytosol) + F
DY
In the absence of ADP, H+ cannot flow
back to the matrix through Fo. The pH and electrical gradients (DpH
& DY) are maximal. As respiration with
outward H+ pumping proceeds, the free energy change for H+
ejection (positive DG) increases and
approaches the magnitude of that for electron transfer (negative
DG). When the coupled reaction becomes
non-spontaneous, respiration stops. This is referred to as a static head. In
fact there is usually a low rate of respiration in the absence of ADP,
attributed to H+ leaks.
When ADP is added, H+ enters the
matrix via Fo, as ATP is synthesized. This reduces the pH and
electrical gradients. DG of H+
ejection decreases. The coupled reaction of electron transfer with H+
ejection becomes spontaneous. Respiration resumes or is stimulated.
|
Uncoupling reagents (uncouplers)
are lipid-soluble weak acids. For example, H+ (shown in red) can
dissociate from the hydroxyl group of the uncoupler
dinitrophenol. Uncouplers
dissolve in the membrane, and function as
carriers for H+.
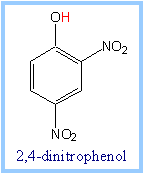
Uncouplers block
oxidative phosphorylation by dissipating the H+ electrochemical
gradient.
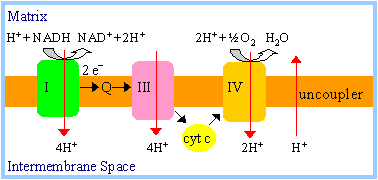
|
|
|
Protons pumped out are
carried by the uncoupler back into the mitochondrial matrix, preventing
development of a pH or electrical gradient. |
|
|
With an uncoupler present there is no pH or
electrical gradient. DG for H+
ejection is zero, and DG for e-
transfer coupled to H+ ejection is maximal (spontaneous).
Respiration proceeds in the presence of an uncoupler, whether or not ADP
is present. Since DG for H+
flux is zero in the absence of a H+ gradient, and hydrolysis of
ATP is spontaneous, the ATP Synthase reaction runs
backward in the presence
of an uncoupler.
An
uncoupling protein (also
called thermogenin) is produced in brown adipose tissue of newborn mammals
and hibernating mammals (see p. 834-835). This protein of the inner
mitochondrial membrane functions as a
H+ carrier.
|
|
The uncoupling protein blocks development of a H+
electrochemical gradient, thereby stimulating respiration. The free energy
change associated with respiration is dissipated as heat. This "non-shivering
thermogenesis" is costly in terms of respiratory energy unavailable for ATP
synthesis, but it provides valuable warming of the organism.
Respiratory chain
inhibitors include the
following:
Rotenone (a common
rat poison) blocks electron transfer in complex I.
Antimycin A blocks
electron transfer in complex III.
Cyanide and
carbon monoxide
inhibit complex IV.
Inhibition at any of these sites will block electron transfer from NADH to
oxygen.
The open axial ligand position of the iron atom
in heme a3 makes it susceptible to binding each of the following
inhibitors: CN-, CO, and the
radical signal molecule ·NO (nitric oxide).
·NO may regulate cellular respiration through its
inhibitory effect, and can induce a condition comparable to hypoxia.
F1FO
ATP SYNTHASE:
|
F1Fo
ATP Synthase of mitochondria, chloroplasts, and
bacteria is represented schematically at right. When the electrochemical H+
gradient is favorable, F1Fo catalyzes ATP synthesis
coupled to spontaneous H+ flux toward the side of the membrane
where F1 protrudes. E.g., in mitochondria, the pH and electrical
gradients drive H+ transport from the intermembrane space to the
matrix compartment.
If no membrane potential
or pH gradient exists to drive the forward reaction, the Keq
favors the reverse reaction, ATP hydrolysis (ATPase activity).
 
|
|
|
Inhibitors of F1Fo,
that block H+ transport coupled to ATP synthesis or hydrolysis,
include:
- oligomycin, an
antibiotic
- DCCD (dicyclohexylcarbodiimide),
a reagent that reacts with carboxyl groups in hydrophobic environments,
forming a covalent adduct.
|
|
|
Viewed by electron
microscopy with negative staining, the ATP synthase appeared as "lollipops"
on the inner mitochondrial membrane, facing the matrix. Higher resolution
cryo-electron microscopy later showed each lollipop to have two stalks.
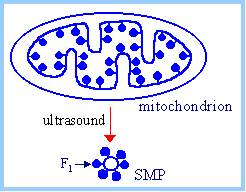
Roles of major subunits
were determined in studies of submitochondrial particles (SMP). If
mitochondria are treated with ultrasound, the inner membrane breaks and
reseals as vesicles, with F1 on the outer surface. Since F1
of intact mitochondria faces the interior matrix space, these SMP are said
to be inside out. |
|
·
F1,
the lollipop head, when extracted from SMP, catalyzes
ATP hydrolysis (the
spontaneous reaction in the absence of an energy input). Thus F1
contains the catalytic domain(s).
·
After removal of F1, the SMP membrane containing Fo
is leaky to H+. Adding back F1 restores the normal low
permeability to H+. Thus it was established that Fo
includes a "proton channel."
·
Either oligomycin or DCCD blocks the H+ leak in
membranes depleted of F1. Thus oligomycin and DCCD inhibit the ATP
Synthase by interacting with Fo.
The subunit composition of the
ATP Synthase was first established for E. coli, which has an operon that
encodes genes for all subunits. Stalk subunits were classified initially as
being part of either F1 or Fo, based on whether they
co-purified with extracted F1.
- F1 subunits,
as originally classified, were named with Greek letters in order of
decreasing molecular weight. They are present in stoichiometry
a3,
b3,
g,
d,
e.
- The
a and
b subunits (513 and 460 amino
acid residues in E. coli), are homologous to one another. Looking down
at the membrane, a &
b subunits alternate around a
ring. (The g subunit is
discussed below.)
- There are three
nucleotide-binding catalytic sites, located at
ab interfaces but predominantly
involving residues of the b
subunits.
- Each
a subunit contains a tightly
bound ATP, but is inactive in catalysis.
- Adenine nucleotides
bind to both a and
b subunits with Mg++.

|
|
- Fo subunits, as
originally classified, were named in Roman letters with decreasing molecular
weight. The stoichiometry of these subunits in the E. coli is a, b2,
c10.
Mammalian mitochondrial F1Fo
is slightly more complex than the bacterial enzyme, with a few additional
subunits. Also, since names were assigned based on apparent molecular weights,
some subunits were given different names in different organisms.
- The bovine
d subunit turned out to be homologous
to the E. coli e subunit.
- The bovine
e subunit is unique.
- A bovine subunit called OSCP
(oligomycin sensitivity conferral protein) is homologous to the E. coli
d subunit.
- The bovine enzyme has
additional subunits d and F6.
There is evidence that the ATP Synthase (F1Fo)
may form a complex with the
adenine nucleotide
translocase (ADP/ATP antiporter) and the
phosphate carrier
(Pi/H+ symporter). This complex has been designated the
ATP Synthasome.














